Search Results for author: Maria A. Rocca
Found 2 papers, 1 papers with code
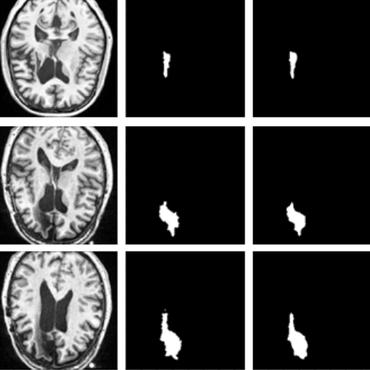
Multi-branch Convolutional Neural Network for Multiple Sclerosis Lesion Segmentation
no code implementations • 7 Nov 2018 • Shahab Aslani, Michael Dayan, Loredana Storelli, Massimo Filippi, Vittorio Murino, Maria A. Rocca, Diego Sona
Our method is based on a deep end-to-end 2D convolutional neural network (CNN) for slice-based segmentation of 3D volumetric data.
Automatic segmentation of the spinal cord and intramedullary multiple sclerosis lesions with convolutional neural networks
2 code implementations • 16 May 2018 • Charley Gros, Benjamin De Leener, Atef Badji, Josefina Maranzano, Dominique Eden, Sara M. Dupont, Jason Talbott, Ren Zhuoquiong, Yaou Liu, Tobias Granberg, Russell Ouellette, Yasuhiko Tachibana, Masaaki Hori, Kouhei Kamiya, Lydia Chougar, Leszek Stawiarz, Jan Hillert, Elise Bannier, Anne Kerbrat, Gilles Edan, Pierre Labauge, Virginie Callot, Jean Pelletier, Bertrand Audoin, Henitsoa Rasoanandrianina, Jean-Christophe Brisset, Paola Valsasina, Maria A. Rocca, Massimo Filippi, Rohit Bakshi, Shahamat Tauhid, Ferran Prados, Marios Yiannakas, Hugh Kearney, Olga Ciccarelli, Seth Smith, Constantina Andrada Treaba, Caterina Mainero, Jennifer Lefeuvre, Daniel S. Reich, Govind Nair, Vincent Auclair, Donald G. McLaren, Allan R. Martin, Michael G. Fehlings, Shahabeddin Vahdat, Ali Khatibi, Julien Doyon, Timothy Shepherd, Erik Charlson, Sridar Narayanan, Julien Cohen-Adad
The goal of this study was to develop a fully-automatic framework, robust to variability in both image parameters and clinical condition, for segmentation of the spinal cord and intramedullary MS lesions from conventional MRI data.