Search Results for author: Tommi Jaakkola
Found 114 papers, 66 papers with code
Predicting deliberative outcomes
no code implementations • ICML 2020 • Vikas Garg, Tommi Jaakkola
Our games take as input, e. g., UN resolution to be voted on, and map such contexts to initial strategies, player utilities, and interactions.
Predicting deliberative outcomes
no code implementations • ICML 2020 • Vikas Garg, Tommi Jaakkola
Our games take as input, e. g., UN resolution to be voted on, and map such contexts to initial strategies, player utilities, and interactions.
Composing Molecules with Multiple Property Constraints
no code implementations • ICML 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
These rationales are identified from molecules as substructures that are likely responsible for each property of interest.
Verlet Flows: Exact-Likelihood Integrators for Flow-Based Generative Models
no code implementations • 5 May 2024 • Ezra Erives, Bowen Jing, Tommi Jaakkola
Approximations in computing model likelihoods with continuous normalizing flows (CNFs) hinder the use of these models for importance sampling of Boltzmann distributions, where exact likelihoods are required.
Deep Confident Steps to New Pockets: Strategies for Docking Generalization
2 code implementations • 28 Feb 2024 • Gabriele Corso, Arthur Deng, Benjamin Fry, Nicholas Polizzi, Regina Barzilay, Tommi Jaakkola
Accurate blind docking has the potential to lead to new biological breakthroughs, but for this promise to be realized, docking methods must generalize well across the proteome.
Dirichlet Flow Matching with Applications to DNA Sequence Design
1 code implementation • 8 Feb 2024 • Hannes Stark, Bowen Jing, Chenyu Wang, Gabriele Corso, Bonnie Berger, Regina Barzilay, Tommi Jaakkola
Further, we provide distilled Dirichlet flow matching, which enables one-step sequence generation with minimal performance hits, resulting in $O(L)$ speedups compared to autoregressive models.
AlphaFold Meets Flow Matching for Generating Protein Ensembles
1 code implementation • 7 Feb 2024 • Bowen Jing, Bonnie Berger, Tommi Jaakkola
When trained and evaluated on the PDB, our method provides a superior combination of precision and diversity compared to AlphaFold with MSA subsampling.
Generative Flows on Discrete State-Spaces: Enabling Multimodal Flows with Applications to Protein Co-Design
1 code implementation • 7 Feb 2024 • Andrew Campbell, Jason Yim, Regina Barzilay, Tom Rainforth, Tommi Jaakkola
Our approach achieves state-of-the-art co-design performance while allowing the same multimodal model to be used for flexible generation of the sequence or structure.
Sample, estimate, aggregate: A recipe for causal discovery foundation models
1 code implementation • 2 Feb 2024 • Menghua Wu, Yujia Bao, Regina Barzilay, Tommi Jaakkola
Causal discovery, the task of inferring causal structure from data, promises to accelerate scientific research, inform policy making, and more.
Correcting Diffusion Generation through Resampling
1 code implementation • 10 Dec 2023 • Yujian Liu, Yang Zhang, Tommi Jaakkola, Shiyu Chang
Despite diffusion models' superior capabilities in modeling complex distributions, there are still non-trivial distributional discrepancies between generated and ground-truth images, which has resulted in several notable problems in image generation, including missing object errors in text-to-image generation and low image quality.
Equivariant Scalar Fields for Molecular Docking with Fast Fourier Transforms
1 code implementation • 7 Dec 2023 • Bowen Jing, Tommi Jaakkola, Bonnie Berger
The runtime of our approach can be amortized at several levels of abstraction, and is particularly favorable for virtual screening settings with a common binding pocket.
Fast non-autoregressive inverse folding with discrete diffusion
1 code implementation • 5 Dec 2023 • John J. Yang, Jason Yim, Regina Barzilay, Tommi Jaakkola
Generating protein sequences that fold into a intended 3D structure is a fundamental step in de novo protein design.
Risk-Controlling Model Selection via Guided Bayesian Optimization
no code implementations • 4 Dec 2023 • Bracha Laufer-Goldshtein, Adam Fisch, Regina Barzilay, Tommi Jaakkola
Adjustable hyperparameters of machine learning models typically impact various key trade-offs such as accuracy, fairness, robustness, or inference cost.
Removing Biases from Molecular Representations via Information Maximization
1 code implementation • 1 Dec 2023 • Chenyu Wang, Sharut Gupta, Caroline Uhler, Tommi Jaakkola
High-throughput drug screening -- using cell imaging or gene expression measurements as readouts of drug effect -- is a critical tool in biotechnology to assess and understand the relationship between the chemical structure and biological activity of a drug.
Learning Interatomic Potentials at Multiple Scales
no code implementations • 20 Oct 2023 • Xiang Fu, Albert Musaelian, Anders Johansson, Tommi Jaakkola, Boris Kozinsky
When running MD, the MTS integrator then evaluates the smaller model for every time step and the larger model less frequently, accelerating simulation.
Particle Guidance: non-I.I.D. Diverse Sampling with Diffusion Models
1 code implementation • 19 Oct 2023 • Gabriele Corso, Yilun Xu, Valentin De Bortoli, Regina Barzilay, Tommi Jaakkola
In light of the widespread success of generative models, a significant amount of research has gone into speeding up their sampling time.
MOFDiff: Coarse-grained Diffusion for Metal-Organic Framework Design
no code implementations • 16 Oct 2023 • Xiang Fu, Tian Xie, Andrew S. Rosen, Tommi Jaakkola, Jake Smith
Metal-organic frameworks (MOFs) are of immense interest in applications such as gas storage and carbon capture due to their exceptional porosity and tunable chemistry.
Harmonic Self-Conditioned Flow Matching for Multi-Ligand Docking and Binding Site Design
1 code implementation • 9 Oct 2023 • Hannes Stärk, Bowen Jing, Regina Barzilay, Tommi Jaakkola
A significant amount of protein function requires binding small molecules, including enzymatic catalysis.
Fast protein backbone generation with SE(3) flow matching
1 code implementation • 8 Oct 2023 • Jason Yim, Andrew Campbell, Andrew Y. K. Foong, Michael Gastegger, José Jiménez-Luna, Sarah Lewis, Victor Garcia Satorras, Bastiaan S. Veeling, Regina Barzilay, Tommi Jaakkola, Frank Noé
We present FrameFlow, a method for fast protein backbone generation using SE(3) flow matching.
Artificial Intelligence for Science in Quantum, Atomistic, and Continuum Systems
1 code implementation • 17 Jul 2023 • Xuan Zhang, Limei Wang, Jacob Helwig, Youzhi Luo, Cong Fu, Yaochen Xie, Meng Liu, Yuchao Lin, Zhao Xu, Keqiang Yan, Keir Adams, Maurice Weiler, Xiner Li, Tianfan Fu, Yucheng Wang, Haiyang Yu, Yuqing Xie, Xiang Fu, Alex Strasser, Shenglong Xu, Yi Liu, Yuanqi Du, Alexandra Saxton, Hongyi Ling, Hannah Lawrence, Hannes Stärk, Shurui Gui, Carl Edwards, Nicholas Gao, Adriana Ladera, Tailin Wu, Elyssa F. Hofgard, Aria Mansouri Tehrani, Rui Wang, Ameya Daigavane, Montgomery Bohde, Jerry Kurtin, Qian Huang, Tuong Phung, Minkai Xu, Chaitanya K. Joshi, Simon V. Mathis, Kamyar Azizzadenesheli, Ada Fang, Alán Aspuru-Guzik, Erik Bekkers, Michael Bronstein, Marinka Zitnik, Anima Anandkumar, Stefano Ermon, Pietro Liò, Rose Yu, Stephan Günnemann, Jure Leskovec, Heng Ji, Jimeng Sun, Regina Barzilay, Tommi Jaakkola, Connor W. Coley, Xiaoning Qian, Xiaofeng Qian, Tess Smidt, Shuiwang Ji
Advances in artificial intelligence (AI) are fueling a new paradigm of discoveries in natural sciences.
Improving Protein Optimization with Smoothed Fitness Landscapes
1 code implementation • 2 Jul 2023 • Andrew Kirjner, Jason Yim, Raman Samusevich, Shahar Bracha, Tommi Jaakkola, Regina Barzilay, Ila Fiete
The ability to engineer novel proteins with higher fitness for a desired property would be revolutionary for biotechnology and medicine.
Restart Sampling for Improving Generative Processes
1 code implementation • NeurIPS 2023 • Yilun Xu, Mingyang Deng, Xiang Cheng, Yonglong Tian, Ziming Liu, Tommi Jaakkola
Restart not only outperforms the previous best SDE results, but also accelerates the sampling speed by 10-fold / 2-fold on CIFAR-10 / ImageNet $64 \times 64$.
Towards Coherent Image Inpainting Using Denoising Diffusion Implicit Models
1 code implementation • 6 Apr 2023 • Guanhua Zhang, Jiabao Ji, Yang Zhang, Mo Yu, Tommi Jaakkola, Shiyu Chang
COPAINT also uses the Bayesian framework to jointly modify both revealed and unrevealed regions, but approximates the posterior distribution in a way that allows the errors to gradually drop to zero throughout the denoising steps, thus strongly penalizing any mismatches with the reference image.
GenPhys: From Physical Processes to Generative Models
no code implementations • 5 Apr 2023 • Ziming Liu, Di Luo, Yilun Xu, Tommi Jaakkola, Max Tegmark
We introduce a general family, Generative Models from Physical Processes (GenPhys), where we translate partial differential equations (PDEs) describing physical processes to generative models.
EigenFold: Generative Protein Structure Prediction with Diffusion Models
1 code implementation • 5 Apr 2023 • Bowen Jing, Ezra Erives, Peter Pao-Huang, Gabriele Corso, Bonnie Berger, Tommi Jaakkola
Protein structure prediction has reached revolutionary levels of accuracy on single structures, yet distributional modeling paradigms are needed to capture the conformational ensembles and flexibility that underlie biological function.
PFGM++: Unlocking the Potential of Physics-Inspired Generative Models
1 code implementation • 8 Feb 2023 • Yilun Xu, Ziming Liu, Yonglong Tian, Shangyuan Tong, Max Tegmark, Tommi Jaakkola
The new models reduce to PFGM when $D{=}1$ and to diffusion models when $D{\to}\infty$.
 Ranked #1 on
Image Generation
on FFHQ 64x64 - 4x upscaling
Ranked #1 on
Image Generation
on FFHQ 64x64 - 4x upscaling
SE(3) diffusion model with application to protein backbone generation
1 code implementation • 5 Feb 2023 • Jason Yim, Brian L. Trippe, Valentin De Bortoli, Emile Mathieu, Arnaud Doucet, Regina Barzilay, Tommi Jaakkola
The design of novel protein structures remains a challenge in protein engineering for applications across biomedicine and chemistry.
Stable Target Field for Reduced Variance Score Estimation in Diffusion Models
1 code implementation • 1 Feb 2023 • Yilun Xu, Shangyuan Tong, Tommi Jaakkola
We show that the procedure indeed helps in the challenging intermediate regime by reducing (the trace of) the covariance of training targets.
 Ranked #12 on
Image Generation
on CIFAR-10
Ranked #12 on
Image Generation
on CIFAR-10
Is Conditional Generative Modeling all you need for Decision-Making?
no code implementations • 28 Nov 2022 • Anurag Ajay, Yilun Du, Abhi Gupta, Joshua Tenenbaum, Tommi Jaakkola, Pulkit Agrawal
We further demonstrate the advantages of modeling policies as conditional diffusion models by considering two other conditioning variables: constraints and skills.
Efficiently Controlling Multiple Risks with Pareto Testing
no code implementations • 14 Oct 2022 • Bracha Laufer-Goldshtein, Adam Fisch, Regina Barzilay, Tommi Jaakkola
Machine learning applications frequently come with multiple diverse objectives and constraints that can change over time.
Forces are not Enough: Benchmark and Critical Evaluation for Machine Learning Force Fields with Molecular Simulations
1 code implementation • 13 Oct 2022 • Xiang Fu, Zhenghao Wu, Wujie Wang, Tian Xie, Sinan Keten, Rafael Gomez-Bombarelli, Tommi Jaakkola
We benchmark a collection of state-of-the-art (SOTA) ML FF models and illustrate, in particular, how the commonly benchmarked force accuracy is not well aligned with relevant simulation metrics.
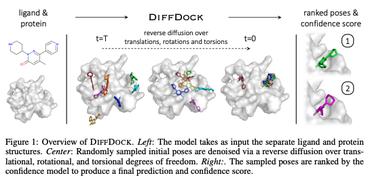
DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking
2 code implementations • 4 Oct 2022 • Gabriele Corso, Hannes Stärk, Bowen Jing, Regina Barzilay, Tommi Jaakkola
We instead frame molecular docking as a generative modeling problem and develop DiffDock, a diffusion generative model over the non-Euclidean manifold of ligand poses.
 Ranked #1 on
Blind Docking
on PDBbind
Ranked #1 on
Blind Docking
on PDBbind
Poisson Flow Generative Models
1 code implementation • 22 Sep 2022 • Yilun Xu, Ziming Liu, Max Tegmark, Tommi Jaakkola
We interpret the data points as electrical charges on the $z=0$ hyperplane in a space augmented with an additional dimension $z$, generating a high-dimensional electric field (the gradient of the solution to Poisson equation).
 Ranked #31 on
Image Generation
on CIFAR-10
Ranked #31 on
Image Generation
on CIFAR-10
Calibrated Selective Classification
no code implementations • 25 Aug 2022 • Adam Fisch, Tommi Jaakkola, Regina Barzilay
Providing calibrated uncertainty estimates alongside predictions -- probabilities that correspond to true frequencies -- can be as important as having predictions that are simply accurate on average.
Antibody-Antigen Docking and Design via Hierarchical Equivariant Refinement
1 code implementation • 14 Jul 2022 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
The binding affinity is governed by the 3D binding interface where antibody residues (paratope) closely interact with antigen residues (epitope).
Diffusion probabilistic modeling of protein backbones in 3D for the motif-scaffolding problem
1 code implementation • 8 Jun 2022 • Brian L. Trippe, Jason Yim, Doug Tischer, David Baker, Tamara Broderick, Regina Barzilay, Tommi Jaakkola
Construction of a scaffold structure that supports a desired motif, conferring protein function, shows promise for the design of vaccines and enzymes.
Torsional Diffusion for Molecular Conformer Generation
1 code implementation • 1 Jun 2022 • Bowen Jing, Gabriele Corso, Jeffrey Chang, Regina Barzilay, Tommi Jaakkola
Molecular conformer generation is a fundamental task in computational chemistry.
Subspace Diffusion Generative Models
1 code implementation • 3 May 2022 • Bowen Jing, Gabriele Corso, Renato Berlinghieri, Tommi Jaakkola
Score-based models generate samples by mapping noise to data (and vice versa) via a high-dimensional diffusion process.
 Ranked #19 on
Image Generation
on CIFAR-10
Ranked #19 on
Image Generation
on CIFAR-10
Simulate Time-integrated Coarse-grained Molecular Dynamics with Multi-Scale Graph Networks
1 code implementation • 21 Apr 2022 • Xiang Fu, Tian Xie, Nathan J. Rebello, Bradley D. Olsen, Tommi Jaakkola
Molecular dynamics (MD) simulation is essential for various scientific domains but computationally expensive.
Conformal Prediction Sets with Limited False Positives
1 code implementation • 15 Feb 2022 • Adam Fisch, Tal Schuster, Tommi Jaakkola, Regina Barzilay
We propose to trade coverage for a notion of precision by enforcing that the presence of incorrect candidates in the predicted conformal sets (i. e., the total number of false positives) is bounded according to a user-specified tolerance.
EquiBind: Geometric Deep Learning for Drug Binding Structure Prediction
1 code implementation • 7 Feb 2022 • Hannes Stärk, Octavian-Eugen Ganea, Lagnajit Pattanaik, Regina Barzilay, Tommi Jaakkola
Predicting how a drug-like molecule binds to a specific protein target is a core problem in drug discovery.
 Ranked #5 on
Blind Docking
on PDBBind
Ranked #5 on
Blind Docking
on PDBBind
Controlling Directions Orthogonal to a Classifier
1 code implementation • ICLR 2022 • Yilun Xu, Hao He, Tianxiao Shen, Tommi Jaakkola
We propose to identify directions invariant to a given classifier so that these directions can be controlled in tasks such as style transfer.
Independent SE(3)-Equivariant Models for End-to-End Rigid Protein Docking
1 code implementation • ICLR 2022 • Octavian-Eugen Ganea, Xinyuan Huang, Charlotte Bunne, Yatao Bian, Regina Barzilay, Tommi Jaakkola, Andreas Krause
Protein complex formation is a central problem in biology, being involved in most of the cell's processes, and essential for applications, e. g. drug design or protein engineering.
Fragment-based Sequential Translation for Molecular Optimization
no code implementations • NeurIPS Workshop AI4Scien 2021 • Benson Chen, Xiang Fu, Regina Barzilay, Tommi Jaakkola
Equipped with the learned fragment vocabulary, we propose Fragment-based Sequential Translation (FaST), which learns a reinforcement learning (RL) policy to iteratively translate model-discovered molecules into increasingly novel molecules while satisfying desired properties.
Learning Representations that Support Robust Transfer of Predictors
2 code implementations • 19 Oct 2021 • Yilun Xu, Tommi Jaakkola
We further demonstrate the impact of optimizing such transfer risk on two controlled settings, each representing a different pattern of environment shift, as well as on two real-world datasets.
Crystal Diffusion Variational Autoencoder for Periodic Material Generation
4 code implementations • ICLR 2022 • Tian Xie, Xiang Fu, Octavian-Eugen Ganea, Regina Barzilay, Tommi Jaakkola
Generating the periodic structure of stable materials is a long-standing challenge for the material design community.
Iterative Refinement Graph Neural Network for Antibody Sequence-Structure Co-design
1 code implementation • ICLR 2022 • Wengong Jin, Jeremy Wohlwend, Regina Barzilay, Tommi Jaakkola
In this paper, we propose a generative model to automatically design the CDRs of antibodies with enhanced binding specificity or neutralization capabilities.
Learning Task Informed Abstractions
1 code implementation • 29 Jun 2021 • Xiang Fu, Ge Yang, Pulkit Agrawal, Tommi Jaakkola
Current model-based reinforcement learning methods struggle when operating from complex visual scenes due to their inability to prioritize task-relevant features.
 Model-based Reinforcement Learning
Model-based Reinforcement Learning
 reinforcement-learning
+1
reinforcement-learning
+1
Mol2Image: Improved Conditional Flow Models for Molecule to Image Synthesis
no code implementations • CVPR 2021 • Karren Yang, Samuel Goldman, Wengong Jin, Alex X. Lu, Regina Barzilay, Tommi Jaakkola, Caroline Uhler
In this paper, we aim to synthesize cell microscopy images under different molecular interventions, motivated by practical applications to drug development.
Consistent Accelerated Inference via Confident Adaptive Transformers
1 code implementation • EMNLP 2021 • Tal Schuster, Adam Fisch, Tommi Jaakkola, Regina Barzilay
In this work, we present CATs -- Confident Adaptive Transformers -- in which we simultaneously increase computational efficiency, while guaranteeing a specifiable degree of consistency with the original model with high confidence.
Few-shot Conformal Prediction with Auxiliary Tasks
1 code implementation • 17 Feb 2021 • Adam Fisch, Tal Schuster, Tommi Jaakkola, Regina Barzilay
We develop a novel approach to conformal prediction when the target task has limited data available for training.
Discovering Synergistic Drug Combinations for COVID with Biological Bottleneck Models
no code implementations • 9 Nov 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
Drug combinations play an important role in therapeutics due to its better efficacy and reduced toxicity.
Information Obfuscation of Graph Neural Networks
1 code implementation • 28 Sep 2020 • Peiyuan Liao, Han Zhao, Keyulu Xu, Tommi Jaakkola, Geoffrey Gordon, Stefanie Jegelka, Ruslan Salakhutdinov
While the advent of Graph Neural Networks (GNNs) has greatly improved node and graph representation learning in many applications, the neighborhood aggregation scheme exposes additional vulnerabilities to adversaries seeking to extract node-level information about sensitive attributes.
Efficient Conformal Prediction via Cascaded Inference with Expanded Admission
1 code implementation • ICLR 2021 • Adam Fisch, Tal Schuster, Tommi Jaakkola, Regina Barzilay
This set is guaranteed to contain a correct answer with high probability, and is well-suited for many open-ended classification tasks.
Improved Conditional Flow Models for Molecule to Image Synthesis
1 code implementation • 15 Jun 2020 • Karren Yang, Samuel Goldman, Wengong Jin, Alex Lu, Regina Barzilay, Tommi Jaakkola, Caroline Uhler
In this paper, we aim to synthesize cell microscopy images under different molecular interventions, motivated by practical applications to drug development.
Optimal Transport Graph Neural Networks
2 code implementations • 8 Jun 2020 • Benson Chen, Gary Bécigneul, Octavian-Eugen Ganea, Regina Barzilay, Tommi Jaakkola
Current graph neural network (GNN) architectures naively average or sum node embeddings into an aggregated graph representation -- potentially losing structural or semantic information.
 Ranked #1 on
Graph Regression
on Lipophilicity
(using extra training data)
Ranked #1 on
Graph Regression
on Lipophilicity
(using extra training data)
Enforcing Predictive Invariance across Structured Biomedical Domains
no code implementations • 6 Jun 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
We evaluate our method on multiple applications: molecular property prediction, protein homology and stability prediction and show that RGM significantly outperforms previous state-of-the-art baselines.
Adaptive Invariance for Molecule Property Prediction
no code implementations • 5 May 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
Effective property prediction methods can help accelerate the search for COVID-19 antivirals either through accurate in-silico screens or by effectively guiding on-going at-scale experimental efforts.
Self-Supervised Learning of Appliance Usage
no code implementations • ICLR 2020 • Chen-Yu Hsu, Abbas Zeitoun, Guang-He Lee, Dina Katabi, Tommi Jaakkola
We show that this cross-modal prediction task allows us to detect when a particular appliance is used, and the location of the appliance in the home, all in a self-supervised manner, without any labeled data.
The Benefits of Pairwise Discriminators for Adversarial Training
1 code implementation • 20 Feb 2020 • Shangyuan Tong, Timur Garipov, Tommi Jaakkola
We provide sufficient conditions for local convergence; characterize the capacity balance that should guide the discriminator and generator choices; and construct examples of minimally sufficient discriminators.
Generalization and Representational Limits of Graph Neural Networks
no code implementations • ICML 2020 • Vikas K. Garg, Stefanie Jegelka, Tommi Jaakkola
We address two fundamental questions about graph neural networks (GNNs).
Improving Molecular Design by Stochastic Iterative Target Augmentation
2 code implementations • ICML 2020 • Kevin Yang, Wengong Jin, Kyle Swanson, Regina Barzilay, Tommi Jaakkola
The property predictor is then used as a likelihood model for filtering candidate structures from the generative model.
Multi-Objective Molecule Generation using Interpretable Substructures
4 code implementations • 8 Feb 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
These rationales are identified from molecules as substructures that are likely responsible for each property of interest.
Blank Language Models
1 code implementation • EMNLP 2020 • Tianxiao Shen, Victor Quach, Regina Barzilay, Tommi Jaakkola
We propose Blank Language Model (BLM), a model that generates sequences by dynamically creating and filling in blanks.
Hierarchical Generation of Molecular Graphs using Structural Motifs
2 code implementations • ICML 2020 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
Indeed, as we demonstrate, their performance degrades significantly for larger molecules.
Generative Models for Graph-Based Protein Design
1 code implementation • ICLR Workshop DeepGenStruct 2019 • John Ingraham, Vikas Garg, Regina Barzilay, Tommi Jaakkola
Engineered proteins offer the potential to solve many problems in biomedicine, energy, and materials science, but creating designs that succeed is difficult in practice.
Direct Optimization through \arg \max for Discrete Variational Auto-Encoder
1 code implementation • NeurIPS 2019 • Guy Lorberbom, Tommi Jaakkola, Andreea Gane, Tamir Hazan
Reparameterization of variational auto-encoders with continuous random variables is an effective method for reducing the variance of their gradient estimates.
Denoising Improves Latent Space Geometry in Text Autoencoders
no code implementations • 25 Sep 2019 • Tianxiao Shen, Jonas Mueller, Regina Barzilay, Tommi Jaakkola
Neural language models have recently shown impressive gains in unconditional text generation, but controllable generation and manipulation of text remain challenging.
Iterative Target Augmentation for Effective Conditional Generation
no code implementations • 25 Sep 2019 • Kevin Yang, Wengong Jin, Kyle Swanson, Regina Barzilay, Tommi Jaakkola
Many challenging prediction problems, from molecular optimization to program synthesis, involve creating complex structured objects as outputs.
Hierarchical Graph-to-Graph Translation for Molecules
1 code implementation • 11 Jun 2019 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
The problem of accelerating drug discovery relies heavily on automatic tools to optimize precursor molecules to afford them with better biochemical properties.
 Ranked #1 on
Drug Discovery
on QED
Ranked #1 on
Drug Discovery
on QED
Solving graph compression via optimal transport
no code implementations • NeurIPS 2019 • Vikas K. Garg, Tommi Jaakkola
The transport problem is seeded with prior information about node importance, attributes, and edges in the graph.
Educating Text Autoencoders: Latent Representation Guidance via Denoising
3 code implementations • ICML 2020 • Tianxiao Shen, Jonas Mueller, Regina Barzilay, Tommi Jaakkola
We prove that this simple modification guides the latent space geometry of the resulting model by encouraging the encoder to map similar texts to similar latent representations.
Path-Augmented Graph Transformer Network
2 code implementations • 29 May 2019 • Benson Chen, Regina Barzilay, Tommi Jaakkola
Much of the recent work on learning molecular representations has been based on Graph Convolution Networks (GCN).
Strategic Prediction with Latent Aggregative Games
no code implementations • 29 May 2019 • Vikas K. Garg, Tommi Jaakkola
We introduce a new class of context dependent, incomplete information games to serve as structured prediction models for settings with significant strategic interactions.
Alignment Based Mathching Networks for One-Shot Classification and Open-Set Recognition
no code implementations • ICLR 2019 • Paresh Malalur, Tommi Jaakkola
An attention mechanism can be used to highlight the area of the image that the model focuses on thus offering a narrow view into the mechanism of classification.
Learning Multimodal Graph-to-Graph Translation for Molecule Optimization
no code implementations • ICLR 2019 • Wengong Jin, Kevin Yang, Regina Barzilay, Tommi Jaakkola
We evaluate our model on multiple molecule optimization tasks and show that our model outperforms previous state-of-the-art baselines by a significant margin.
Analyzing Learned Molecular Representations for Property Prediction
4 code implementations • 2 Apr 2019 • Kevin Yang, Kyle Swanson, Wengong Jin, Connor Coley, Philipp Eiden, Hua Gao, Angel Guzman-Perez, Timothy Hopper, Brian Kelley, Miriam Mathea, Andrew Palmer, Volker Settels, Tommi Jaakkola, Klavs Jensen, Regina Barzilay
In addition, we introduce a graph convolutional model that consistently matches or outperforms models using fixed molecular descriptors as well as previous graph neural architectures on both public and proprietary datasets.
 Ranked #3 on
Molecular Property Prediction
on QM9
Ranked #3 on
Molecular Property Prediction
on QM9
Alignment Based Matching Networks for One-Shot Classification and Open-Set Recognition
no code implementations • 11 Mar 2019 • Paresh Malalur, Tommi Jaakkola
An attention mechanism can be used to highlight the area of the image that the model focuses on thus offering a narrow view into the mechanism of classification.
Learning Multimodal Graph-to-Graph Translation for Molecular Optimization
5 code implementations • 3 Dec 2018 • Wengong Jin, Kevin Yang, Regina Barzilay, Tommi Jaakkola
We evaluate our model on multiple molecular optimization tasks and show that our model outperforms previous state-of-the-art baselines.
Towards Robust Interpretability with Self-Explaining Neural Networks
2 code implementations • NeurIPS 2018 • David Alvarez Melis, Tommi Jaakkola
Most recent work on interpretability of complex machine learning models has focused on estimating a-posteriori explanations for previously trained models around specific predictions.
The Variational Homoencoder: Learning to learn high capacity generative models from few examples
1 code implementation • 24 Jul 2018 • Luke B. Hewitt, Maxwell I. Nye, Andreea Gane, Tommi Jaakkola, Joshua B. Tenenbaum
However, when this generative model is expressed as a powerful neural network such as a PixelCNN, we show that existing learning techniques typically fail to effectively use latent variables.
Direct Optimization through $\arg \max$ for Discrete Variational Auto-Encoder
2 code implementations • ICLR 2019 • Guy Lorberbom, Andreea Gane, Tommi Jaakkola, Tamir Hazan
We demonstrate empirically the effectiveness of the direct loss minimization technique in variational autoencoders with both unstructured and structured discrete latent variables.
Junction Tree Variational Autoencoder for Molecular Graph Generation
11 code implementations • ICML 2018 • Wengong Jin, Regina Barzilay, Tommi Jaakkola
We evaluate our model on multiple tasks ranging from molecular generation to optimization.
 Ranked #1 on
Molecular Graph Generation
on InterBioScreen
Ranked #1 on
Molecular Graph Generation
on InterBioScreen
Using Deep Reinforcement Learning to Generate Rationales for Molecules
no code implementations • ICLR 2018 • Benson Chen, Connor Coley, Regina Barzilay, Tommi Jaakkola
Deep learning algorithms are increasingly used in modeling chemical processes.
The Variational Homoencoder: Learning to Infer High-Capacity Generative Models from Few Examples
no code implementations • ICLR 2018 • Luke Hewitt, Andrea Gane, Tommi Jaakkola, Joshua B. Tenenbaum
Hierarchical Bayesian methods have the potential to unify many related tasks (e. g. k-shot classification, conditional, and unconditional generation) by framing each as inference within a single generative model.
Local Aggregative Games
no code implementations • NeurIPS 2017 • Vikas Garg, Tommi Jaakkola
Aggregative games provide a rich abstraction to model strategic multi-agent interactions.
Predicting Organic Reaction Outcomes with Weisfeiler-Lehman Network
1 code implementation • NeurIPS 2017 • Wengong Jin, Connor W. Coley, Regina Barzilay, Tommi Jaakkola
The prediction of organic reaction outcomes is a fundamental problem in computational chemistry.
Sequence to Better Sequence: Continuous Revision of Combinatorial Structures
no code implementations • ICML 2017 • Jonas Mueller, David Gifford, Tommi Jaakkola
Under this model, gradient methods can be used to efficiently optimize the continuous latent factors with respect to inferred outcomes.
Grounding Language for Transfer in Deep Reinforcement Learning
1 code implementation • 1 Aug 2017 • Karthik Narasimhan, Regina Barzilay, Tommi Jaakkola
In this paper, we explore the utilization of natural language to drive transfer for reinforcement learning (RL).
Style Transfer from Non-Parallel Text by Cross-Alignment
12 code implementations • NeurIPS 2017 • Tianxiao Shen, Tao Lei, Regina Barzilay, Tommi Jaakkola
We demonstrate the effectiveness of this cross-alignment method on three tasks: sentiment modification, decipherment of word substitution ciphers, and recovery of word order.
 Ranked #7 on
Text Style Transfer
on Yelp Review Dataset (Small)
Ranked #7 on
Text Style Transfer
on Yelp Review Dataset (Small)
Deriving Neural Architectures from Sequence and Graph Kernels
no code implementations • ICML 2017 • Tao Lei, Wengong Jin, Regina Barzilay, Tommi Jaakkola
The design of neural architectures for structured objects is typically guided by experimental insights rather than a formal process.
Aspect-augmented Adversarial Networks for Domain Adaptation
1 code implementation • TACL 2017 • Yuan Zhang, Regina Barzilay, Tommi Jaakkola
We introduce a neural method for transfer learning between two (source and target) classification tasks or aspects over the same domain.
Learning Tree Structured Potential Games
no code implementations • NeurIPS 2016 • Vikas Garg, Tommi Jaakkola
Many real phenomena, including behaviors, involve strategic interactions that can be learned from data.
Learning Optimal Interventions
no code implementations • 16 Jun 2016 • Jonas Mueller, David N. Reshef, George Du, Tommi Jaakkola
Assuming the underlying relationship remains invariant under intervention, we develop efficient algorithms to identify the optimal intervention policy from limited data and provide theoretical guarantees for our approach in a Gaussian Process setting.
Rationalizing Neural Predictions
3 code implementations • EMNLP 2016 • Tao Lei, Regina Barzilay, Tommi Jaakkola
Our approach combines two modular components, generator and encoder, which are trained to operate well together.
High Dimensional Inference with Random Maximum A-Posteriori Perturbations
no code implementations • 10 Feb 2016 • Tamir Hazan, Francesco Orabona, Anand D. Sarwate, Subhransu Maji, Tommi Jaakkola
This paper shows that the expected value of perturb-max inference with low dimensional perturbations can be used sequentially to generate unbiased samples from the Gibbs distribution.
Semi-supervised Question Retrieval with Gated Convolutions
1 code implementation • NAACL 2016 • Tao Lei, Hrishikesh Joshi, Regina Barzilay, Tommi Jaakkola, Katerina Tymoshenko, Alessandro Moschitti, Lluis Marquez
Question answering forums are rapidly growing in size with no effective automated ability to refer to and reuse answers already available for previous posted questions.
Principal Differences Analysis: Interpretable Characterization of Differences between Distributions
no code implementations • NeurIPS 2015 • Jonas Mueller, Tommi Jaakkola
We introduce principal differences analysis (PDA) for analyzing differences between high-dimensional distributions.
Steps Toward Deep Kernel Methods from Infinite Neural Networks
no code implementations • 20 Aug 2015 • Tamir Hazan, Tommi Jaakkola
Contemporary deep neural networks exhibit impressive results on practical problems.
Molding CNNs for text: non-linear, non-consecutive convolutions
2 code implementations • EMNLP 2015 • Tao Lei, Regina Barzilay, Tommi Jaakkola
Moreover, we extend the n-gram convolution to non-consecutive words to recognize patterns with intervening words.
Structured Prediction: From Gaussian Perturbations to Linear-Time Principled Algorithms
no code implementations • 5 Aug 2015 • Jean Honorio, Tommi Jaakkola
Thus, using the maximum loss over random structured outputs is a principled way of learning the parameter of structured prediction models.
CRAFT: ClusteR-specific Assorted Feature selecTion
no code implementations • 25 Jun 2015 • Vikas K. Garg, Cynthia Rudin, Tommi Jaakkola
We present a framework for clustering with cluster-specific feature selection.
An Unsupervised Method for Uncovering Morphological Chains
1 code implementation • TACL 2015 • Karthik Narasimhan, Regina Barzilay, Tommi Jaakkola
In contrast, we propose a model for unsupervised morphological analysis that integrates orthographic and semantic views of words.
Controlling privacy in recommender systems
no code implementations • NeurIPS 2014 • Yu Xin, Tommi Jaakkola
Recommender systems involve an inherent trade-off between accuracy of recommendations and the extent to which users are willing to release information about their preferences.
Learning Efficient Random Maximum A-Posteriori Predictors with Non-Decomposable Loss Functions
no code implementations • NeurIPS 2013 • Tamir Hazan, Subhransu Maji, Joseph Keshet, Tommi Jaakkola
In this work we develop efficient methods for learning random MAP predictors for structured label problems.
On Measure Concentration of Random Maximum A-Posteriori Perturbations
no code implementations • 15 Oct 2013 • Francesco Orabona, Tamir Hazan, Anand D. Sarwate, Tommi Jaakkola
Applying the general result to MAP perturbations can yield a more efficient algorithm to approximate sampling from the Gibbs distribution.
On Sampling from the Gibbs Distribution with Random Maximum A-Posteriori Perturbations
no code implementations • NeurIPS 2013 • Tamir Hazan, Subhransu Maji, Tommi Jaakkola
In this paper we describe how MAP inference can be used to sample efficiently from Gibbs distributions.
Proceedings of the Twenty-First Conference on Uncertainty in Artificial Intelligence (2005)
no code implementations • 25 Aug 2012 • Fahiem Bacchus, Tommi Jaakkola
This is the Proceedings of the Twenty-First Conference on Uncertainty in Artificial Intelligence, which was held in Edinburgh, Scotland July 26 - 29 2005.
On the Statistical Efficiency of $\ell_{1,p}$ Multi-Task Learning of Gaussian Graphical Models
no code implementations • 18 Jul 2012 • Jean Honorio, Tommi Jaakkola, Dimitris Samaras
In this paper, we present $\ell_{1, p}$ multi-task structure learning for Gaussian graphical models.